The Conversation
The Conversation
Published: July
We were recently invited to write about our work on active micromachines. The article can be read for free on the Convesation website.
The Conversation
Published: July
We were recently invited to write about our work on active micromachines. The article can be read for free on the Convesation website.
Most of my projects investigate some microscopic object (or objects) such as polymers or microbial swimmers immersed in a simple background fluid. Since it’s the dynamics of the objects that we’re typically interested in, we treat the fluid as simply as possible. In order to do this, we often use the MPCD method (described here).
However, not all fluids are simple. Some biological fluids have intrinsic elasticity, while others are made of or include rod-like molecules. These fluids are called “nematic” — from the Greek word for “threadlike”. Since the molecules are rod-like, they can align; and when they align they are partially ordered. Ordered solids are called crystals and so these fluids are called liquid-crystals. Since many biological molecules are rod-like, such as filamentous-actin, their solutions can be liquid crystalline.
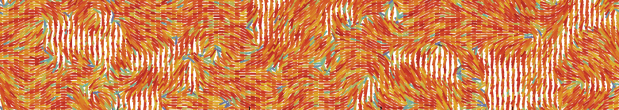

Dealing with flowing liquid crystals is notoriously difficult. When the liquid crystal is just the background fluid to the dynamics of the colloids, polymers, or microbes, significant computational effort must be spent to simulate the background. This is why we developed a computationally efficient and versatile multi-particle collision dynamics (MPCD)-based algorithm to simulate nematohydrodynamic baths.
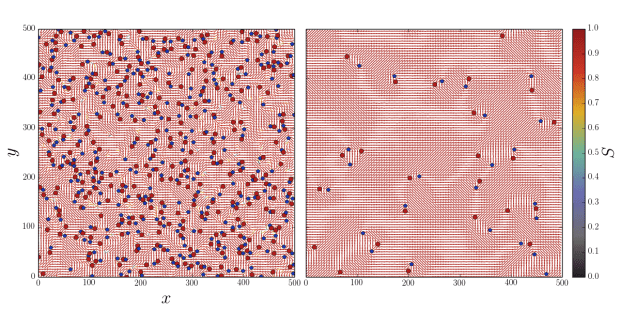
Our key realization was that the MPCD algorithm isn’t limited to solving the Navier-Stokes equation for momentum transport in a fluid, but can really solve any transport equation so long as a suitable “collision” operation (now more generally referred to as an “exchange”) can be defined and the continuum field of interest can be discretized as a particle property. We gave each point-like MPCD particle an orientation that would interact by an order-collision operation based on Maier-Saupe theory for liquid crystals in equilibrium.
This simple algorithm works very well. It is accurate in 2D and 3D; it reproduces defect dynamics; it has an intrinsic elasticity; it can have tumbling or shear aligning regimes when it flows; and it can handle complex boundary conditions and mobile macromolecules, colloids and swimmers.
Microorganisms live in a very different world than us — a world commonly submerged in water. But, at such microscopic sizes, swimming in water seems far more like swimming through tar. Understanding the dynamics of how microorganisms travel through liquids can help us control their spread.
Since the microbial world is a liquid one, flows play an important role in setting microbe motion. Bacteria, for instance, commonly have “run-tumble” dynamics: They swim forward for a time then briefly tumble in a new, random direction. Walls and obstacles can impact these dynamics, not only by simply getting in the way but also by changing the flows caused by the bacteria. In this way, even distant objects interact with the swimmers through flow.
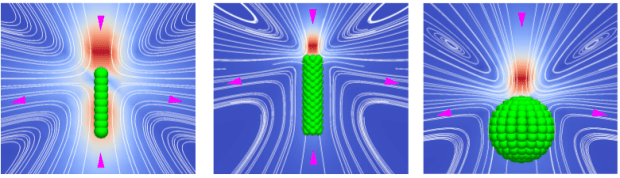
These hydrodynamic interactions are challenging to predict. We’ve worked with Joost de Graaf (University of Edinburgh), who has developed versatile models for computationally simulating the dynamics of self-propelled particles (such as living bacteria) within fluids. These simulations build swimmers out of beads, which appear raspberry-like in shape. The resulting models reproduce well the hydrodynamics of swimming cells, like E. coli or sperm, which means that in the future these models may be able to accurately predict swimming dynamics in complex geometries and flows.
Living near interfaces can be beneficial to microbes. Liquid-air interfaces may be the source of oxygen and sunlight. Solid surfaces accumulate sediments, which can include valuable nutrients. They can also offer anchoring points for the formation of protective biofilm. Films of liquid over a solid wall have both of these types of beneficial interfaces. However, an unavoidable consequence of being near surfaces is a shearing when the film flows. These flows can have a large impact on the swimming dynamics of microbes.
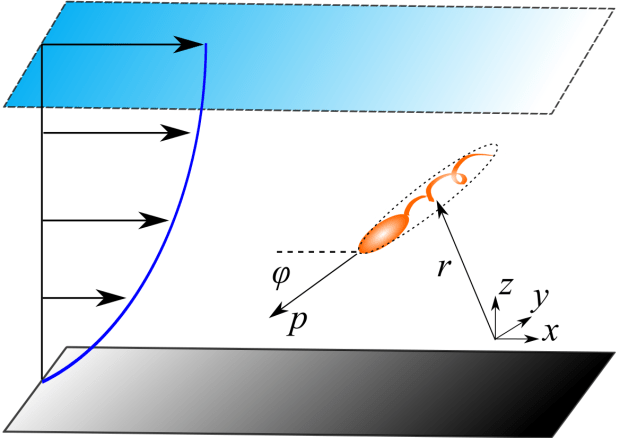
In a series of papers with Arnold Mathijssen and Amin Doostmohammadi, we investigated the swimming dynamics of motile microbes in flowing films. We began by building an analytical model for the flows produced when microbes swim in a film. We were able to reproduce many of the different flows generated by swimmers. These include flagellated bacteria that swim by propelling fluid back and then pushing fluid forward (called “pusher-type swimmers”), and puller-type swimmers that pull fluid toward their head and drag their body forward. We also reproduced ciliated swimmers with their many hair-like flagella.
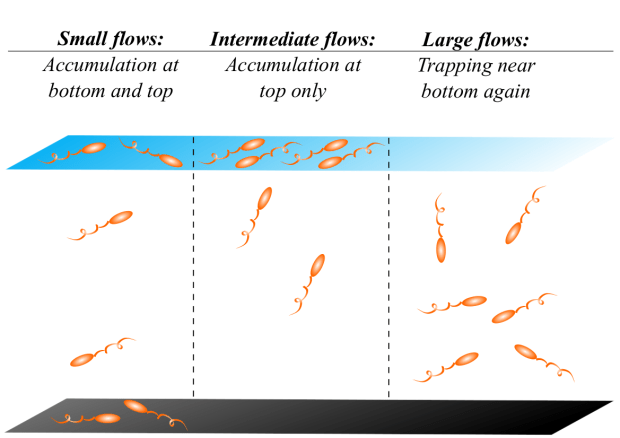
We then put our swimmer models in a flowing film and looked at the dynamics. We found there is a sweet spot for rinsing bacteria away. When the film isn’t flowing, the microbes are attracted most strongly to the solid surface and they stay near the floor until the flow is strong enough to peel microbial swimmers off the surface. That is to say, at moderate flows bacteria in a film are washed away but that’s not the end of it. If much stronger flows are applied, the flow turns the microbes back down towards the surface, where they are more likely to form biofilms.
Run-tumble Dynamics vs. Hydrodynamic Interactions in Films
Swimmers like E. coli have “run-tumble” dynamics, meaning they swim forward for a time (run stage) then rapidly and randomly change to new direction (tumble stage). While they swim in the run stage, they perturb the surrounding fluid and interact with their surroundings through the flow they generate. This causes swimmers to be attracted to walls.
We asked “When does the hydrodynamic attraction win over the randomizing run-tumble dynamics in a film?” We found if we modelled the dynamics as just an effective constant noise then the swimming strategy of the microbes makes quite a difference. However, if the swimmers have a proper run stage followed by a brief and random tumble stage then the swimming strategy is not nearly as important.
We believe this will be of interest to other scientists in our field because it means that when they make simulations of swimmers they should prioritize getting the run-tumble dynamics correct before attempting to reproduce the more complicated but less important hydrodynamics of microbial swimming.
We’ve worked on computational and theoretical models for swimming microbes and looked at their dynamics in biologically relevant environments, such as flowing films. Another important consideration is the nature of the fluid that the microbes swim through. An important aspect of many fluids is that they have a dual fluidic and elastic (viscoelastic) nature. Biological examples of such so-called non-Newtonian fluids include cervical mucus, extracellular matrix surrounding biofilms, and flowing blood. To make matters more complicated, microbes such as sperm and other swimming cells must often swim upstream.
With Arnold Mathijssen and Amin Doostmohammadi, we considered a microswimmer swimming in a channel geometry and subjected to a flow. Previous studies with this set-up were done by Zottl and Stark, who established that the mathematics describing the microbe’s swimming are related to the swinging of a pendulum. We built on their work to predict that swimming cells in biological flows naturally reorient toward the centre of channels, where they migrate upstream. This is because of the elasticity in such fluids.
The reorientation also happens because the flow turns the swimmers away from the walls, like in normal, Newtonian fluids (i.e. water). We found that swimmers in fluids that become more viscous when sheared (like cornstarch that thickens when stirred) swim upstream more quickly than in fluids that thin (like ketchup that spills suddenly after slowly creeping). The swimmers move upstream faster in shear-thickening flows because they can spend more time in slower flows near walls, where they have the chance to sneak further upstream.
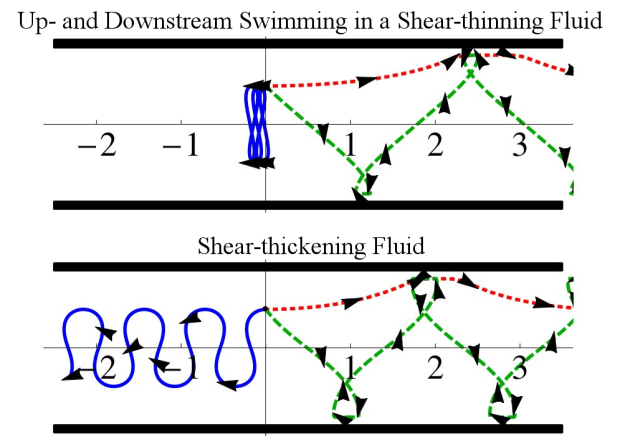
Swimming speed is used to assess the fertility of human semen so we suspect that fertility estimates based on speed could be improved by performing these measurements in viscoelastic fluids. The sperm in such a flowing fluid would swim upstream along the centreline where they can have a fair race, without some cheating by sneaking up along the walls.
This work will soon appear in the journal Physical Review Letters.
Field-Flow Fractionation (FFF; introduced here) separates samples of particles in a flow by applying an external field that causes distinct solutes to have different concentration distributions. The greater the difference in the distributions, the greater the separation of different solutes.
Traditionally, FFF strives to have just a single external force applied to the solutes — more than one external field has been assumed to (a) just increase the net forces when the fields align (termed additive-mode FFF by Bruce Gale) or (b) simply decrease the net forces when the fields compete. One would never apply opposing forces on purpose.

However, while trying to develop models for understanding hyperlayer-FFF (when unavoidable lift forces oppose the external field), I was surprised to find that two opposing fields can produce a dramatic drop in elution time of very specific solute sizes. Gary Slater and I termed this novel technique “adverse-mode FFF”. By applying two strongly-opposing external forces, which scale differently with solute size, a sharp peak occurs in the speed of a solute for a given size. For solutes smaller than the peak position, the force that scales less strongly with size dominates. For larger solutes, the force that scales more strongly with size dominates. Exactly at the peak position, the two opposing forces balance. By increasing both field strengths together, the peak width can be narrowed and be made quite sharp.
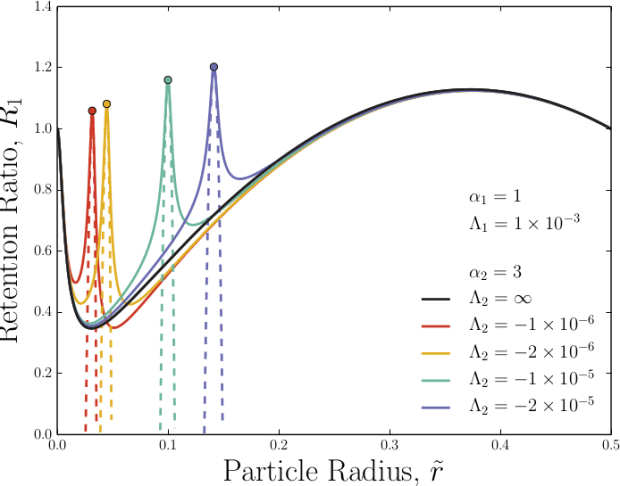
All of this occurs for realizable field strengths. To measure an unknown solute size, adverse-mode FFF could following this recipe:
An external field that scales directly with solute radius (such as flow-FFF, electrical-FFF or thermal-FFF) is applied to an eluting sample.
A second field that scales with the solute volume (radius cubed; such as centrifugal-FFF) is applied to oppose the first field.
By incrementally decreasing the strength of the second field from an initial large value, the peak position would shift to smaller values and an abrupt increase in the speed of the solute would occur when the peak location matches the solute’s size.
The solute size can then be calculated.
Having found the solute size, the accuracy can be improved by increasing both external field strengths together.
DNA doesn’t just float around. It it tugged on and wrapped up by the protein machinary within cells. All of this pushing and pulling occurs within the confined space of the nucleus. In artificial systems, experimentalists pull on DNA and other semi-flexible biomolecules, stretching them within channels or slits.
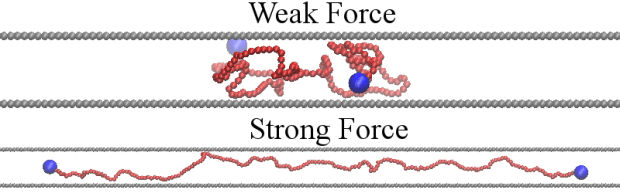
However, the model relating the extension of DNA to the amount of force applied is for fully 3D or perfectly 2D, like in a film. In strongly confining slits, the walls make it easier to stretch the DNA but not as easy as in 2D. The confining walls make it easier because they obstruct the DNA and so limit the number of conformational states that the DNA can have. Since the number of states sets the entropy and the entropy works against the applied stretching force, confinement makes stretching easier.
With Hendrick de Haan (University of Ontario Institute of Technology), we developed a theory for describing the extension of DNA in a channel of any height as a function of force. This theory is built on the classical theory of Marco-Siggia for DNA in 2D and 3D and uses an effective dimensionality between 2 or 3 when the forces are not too strong. When the forces are very strong a mathematical approximation must be applied for strong and weak confinements.
Entropy can be tricky to understand, but one rule often quoted in pop-culture is that entropy increases as time goes on, which leads to more and more disorder. Depletion forces can therefore be surprising since they seem to break this rule-of-thumb; the depletion force is an entropically induced attraction between large colloids due many smaller surrounding “depletant” particles. By bringing the large colloids together, the attraction orders the larger colloids but this is at the benefit of the many small depletants, which gain increased entropy and are more disordered.
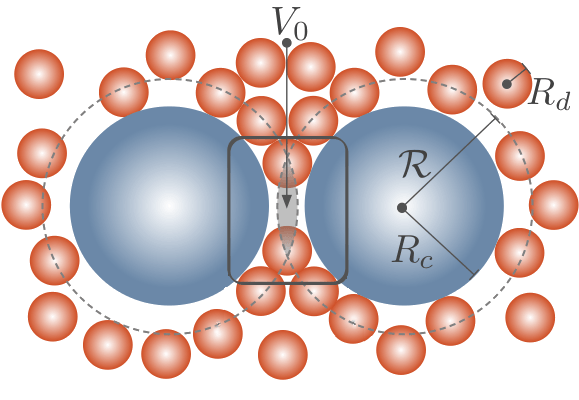
Such entropic forces exist in biological systems. For example, depletion forces lead to fibre bundling, aggregation of red blood cells, and change the reaction kinetics of polymerase chain reactions. They are also ubiquitous in the complex and crowded interior of cells.
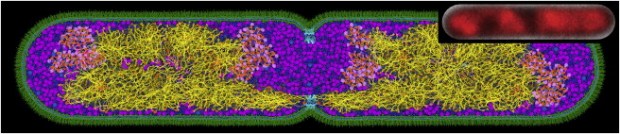
Prior to our work, it was hypothesized that depletion forces acting on chromosomes in simple bacteria by surrounding cytoplasmic proteins could explain the collapse from a swollen conformation to a compact state. We performed simulations to explicitly test whether depletion-induced attraction is actually enough to collapse model chromosomes. The resulting simulations gave us strong evidence that depletion effects are enough to collapse chromosomes; however, this collapse is a continuous phase transition rather than a discontinuous one (as is seen in experiments). This suggests that some proteins must act in specific biological ways (rather than as generic depletants) to fully account for the compaction of nucleoids.
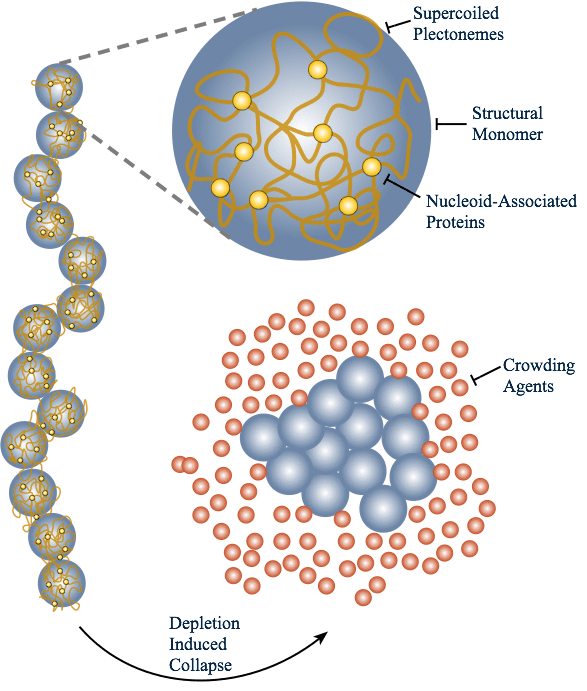
To read more, see our paper on the entropic collapse of coarse-grained chromosomes, or a News and Views by Suckjoon Jun discussing our work.
Mike Foster and Tabaret magazine covered my research and science writing last week. The article can be read on the Tabaret website. The pull out quote that they picked is
The 2014 report by the BSA Media Fellows is now out. Follow the link to read about the fellows’ experiences in some of the best news rooms in the country.
My fellow Fellows were:
You must be logged in to post a comment.