DNA doesn’t just float around. It it tugged on and wrapped up by the protein machinary within cells. All of this pushing and pulling occurs within the confined space of the nucleus. In artificial systems, experimentalists pull on DNA and other semi-flexible biomolecules, stretching them within channels or slits.
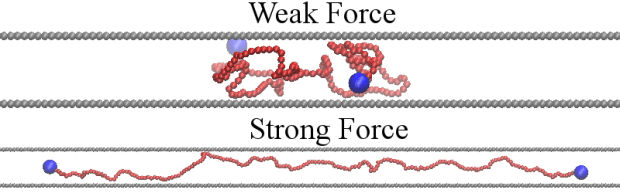
However, the model relating the extension of DNA to the amount of force applied is for fully 3D or perfectly 2D, like in a film. In strongly confining slits, the walls make it easier to stretch the DNA but not as easy as in 2D. The confining walls make it easier because they obstruct the DNA and so limit the number of conformational states that the DNA can have. Since the number of states sets the entropy and the entropy works against the applied stretching force, confinement makes stretching easier.
With Hendrick de Haan (University of Ontario Institute of Technology), we developed a theory for describing the extension of DNA in a channel of any height as a function of force. This theory is built on the classical theory of Marco-Siggia for DNA in 2D and 3D and uses an effective dimensionality between 2 or 3 when the forces are not too strong. When the forces are very strong a mathematical approximation must be applied for strong and weak confinements.

You must be logged in to post a comment.